The Rumbaugh Lab
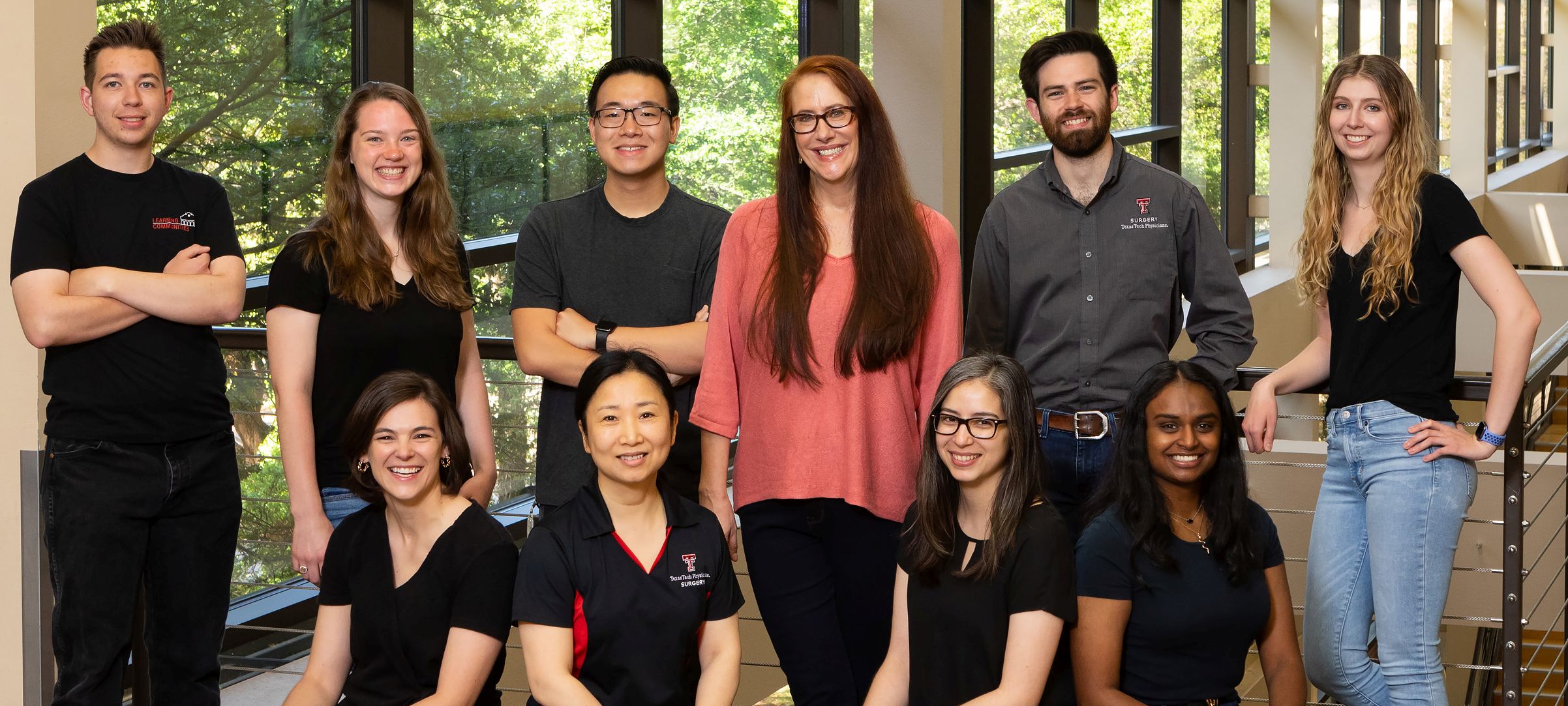

Got Questions?
We're here to help. Contact us if you have questions.
Polymicrobial interactions and Biofilm Formation in Chronic Wounds
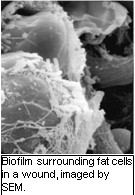 The impact of wound infections on our healthcare system is enormous. In fact, chronically-infected
diabetic foot ulcers are considered the most significant wound care problem in the
world. One major reason that these infections are so difficult to treat is because
bacteria develop ‘biofilms’ after they colonize wounds. A biofilm is a slimy, sugar-rich
structure that encases bacteria and protects them from the immune response and antimicrobials,
making them up to 1,000 times less susceptible to killing. The microbial populations
of chronic wounds are complex and can include dozens of major species, yet very
The impact of wound infections on our healthcare system is enormous. In fact, chronically-infected
diabetic foot ulcers are considered the most significant wound care problem in the
world. One major reason that these infections are so difficult to treat is because
bacteria develop ‘biofilms’ after they colonize wounds. A biofilm is a slimy, sugar-rich
structure that encases bacteria and protects them from the immune response and antimicrobials,
making them up to 1,000 times less susceptible to killing. The microbial populations
of chronic wounds are complex and can include dozens of major species, yet very 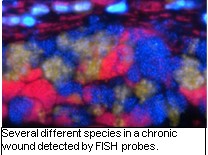 little is understood how interactions between these species influence the course of
infection or affect therapeutic success. We have developed polymicrobial wound models
in order to investigate how different species in wounds work together to build better
biofilms and create more difficult to treat infections.
little is understood how interactions between these species influence the course of
infection or affect therapeutic success. We have developed polymicrobial wound models
in order to investigate how different species in wounds work together to build better
biofilms and create more difficult to treat infections.
Pseudomonas aeruginosa Pathogenesis in Wounds
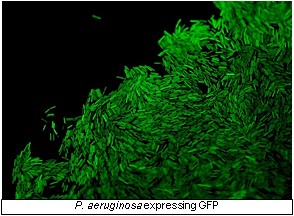 Infection with the Gram-negative pathogen Pseudomonas aeruginosa is one of the major
causes of morbidity and mortality in severely burned patients, and the cause of debilitating
chronic infections in diabetic patients. P. aeruginosa relies on an arsenal of cell-associated
and secreted virulence factors to colonize and infect its host, and it persists and
invades the immune system by building biofilms.
Infection with the Gram-negative pathogen Pseudomonas aeruginosa is one of the major
causes of morbidity and mortality in severely burned patients, and the cause of debilitating
chronic infections in diabetic patients. P. aeruginosa relies on an arsenal of cell-associated
and secreted virulence factors to colonize and infect its host, and it persists and
invades the immune system by building biofilms.
 In collaboration with Marvin Whiteley’s group, http://whiteleylab.biosci.gatech.edu/?q=home, we are using advanced genomic approaches to understand the pathogenesis of P. aeruginosa
in human wounds and mouse models that recapitulate disease.
In collaboration with Marvin Whiteley’s group, http://whiteleylab.biosci.gatech.edu/?q=home, we are using advanced genomic approaches to understand the pathogenesis of P. aeruginosa
in human wounds and mouse models that recapitulate disease.
Necrotizing Soft Tissue Infections
Necrotizing soft tissue infections (NSTIs) are a group of infections that are rapidly progressive and require immediate surgical intervention. The pathogenesis and microbiology of these infections are understudied, which partially explains the relatively stagnant and high mortality despite aggressive surgical and medical treatments. We have recently shown through human metagenomics studies that obligate anaerobes are abundant in NSTIs and are associated with a worsened prognosis. Currently, we are focused on better understanding the polymicrobial interactions behind NSTIs through the study of model organisms, Staphylococcus aureus and Bacteroides fragilis. We hypothesize that microbial interactions that contain obligate anaerobes cause a worsened NSTI compared to those with facultative anaerobes alone.
Development of Biofilm Degrading Agents
Biofilms are communities of microorganisms protected by a thick, self-synthesized extracellular matrix of polysaccharides, proteins, lipids and extracellular DNA. As much as 85% of all bacterial infections are biofilm-associated, compounding the problem of rising antibiotic resistance by offering the resident microbes greatly increased tolerances to antimicrobials. We are tackling this problem by developing agents that degrade the protective extracellular matrix, thereby augmenting the effectiveness of existing antimicrobial agents. We have shown that biofilm polysaccharides can be degraded by enzymes that hydrolyze the glycosidic bonds between the individual sugar moieties, thereby causing the structural collapse of the biofilms, and potentiating the action of antibiotics against the newly liberated bacterial cells in vivo. Currently, we are working on optimizing enzymatic to break down the biofilms present in complex, polymicrobial, clinical infections in a broad-spectrum manner. In doing so, we hope to be able to create a widely-applicable topical treatment that can greatly increase the effectiveness of existing antibiotics, reversing some of the devastating effects of antibiotic resistance on the healthcare field.
Preclinical Efficacy Testing of Experimental Antimicrobials
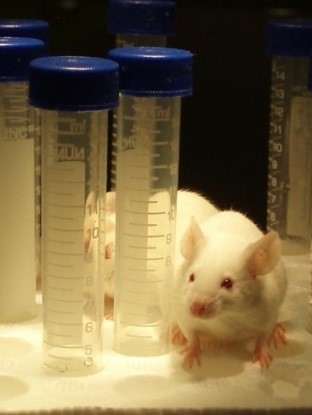 We have developed several in vitro and in vivo wound models with which to test experimental
antimicrobials and anti-biofilm agents. We have extensive experience in working with
companies and academic collaborators to test the efficacy of new therapeutics.
We have developed several in vitro and in vivo wound models with which to test experimental
antimicrobials and anti-biofilm agents. We have extensive experience in working with
companies and academic collaborators to test the efficacy of new therapeutics.
Interkingdom signaling between P. aeruginosa Quorum Sensing Molecules and Host Cells
 Quorum sensing (QS) is a cell density-dependent signaling process used by many bacteria
to coordinate gene expression in a population. QS in Gram-negative bacteria is controlled
by diffusible molecules called autoinducers (AI) that function as ligands for regulatable
transcription factors. At least two separate QS systems exist in P. aeruginosa, the
LasI/LasR and RhlI/RhlR systems. The ligands for LasR and RhlR are N-3-oxododecanoyl-
and N-butyryl- homoserine lactones, or PAI-1 and PAI-2, respectively. Several studies
indicate that bacterial autoinducers, and PAI-1 in particular, can also influence
gene expression in host eukaryotic cells, a process we’ve termed interkingdom signaling.
We hypothesize that this regulatory process involves autoinducer receptor
Quorum sensing (QS) is a cell density-dependent signaling process used by many bacteria
to coordinate gene expression in a population. QS in Gram-negative bacteria is controlled
by diffusible molecules called autoinducers (AI) that function as ligands for regulatable
transcription factors. At least two separate QS systems exist in P. aeruginosa, the
LasI/LasR and RhlI/RhlR systems. The ligands for LasR and RhlR are N-3-oxododecanoyl-
and N-butyryl- homoserine lactones, or PAI-1 and PAI-2, respectively. Several studies
indicate that bacterial autoinducers, and PAI-1 in particular, can also influence
gene expression in host eukaryotic cells, a process we’ve termed interkingdom signaling.
We hypothesize that this regulatory process involves autoinducer receptor  molecules in the host cells, possibly transcription factors. We have shown that P.
aeruginosa autoinducers can efficiently enter mammalian cells and modulate gene expression
potentially through the interaction of nuclear hormone receptors. In addition, we
have also recently shown that the nematode C. elegans can sense bacterial autoinducers
and use this sensory information to ‘learn’ to avoid pathogens.
molecules in the host cells, possibly transcription factors. We have shown that P.
aeruginosa autoinducers can efficiently enter mammalian cells and modulate gene expression
potentially through the interaction of nuclear hormone receptors. In addition, we
have also recently shown that the nematode C. elegans can sense bacterial autoinducers
and use this sensory information to ‘learn’ to avoid pathogens.
Quorum Sensing: Methods and Protocols
 Since its early days in the 1990s, the Quorum Sensing (QS) field has grown from a
few dozen laboratories, investigating the pathways, proteins, and chemicals that facilitate
signaling in bacteria, to hundreds of groups that have integrated evolutionary biology,
computer science, mathematics, engineering, and metagenomics to create an ever-expanding
and dynamic field. In Quorum Sensing: Methods and Protocols, expert researchers provide
an in-depth set of diverse protocols that span this broad area of study.
Since its early days in the 1990s, the Quorum Sensing (QS) field has grown from a
few dozen laboratories, investigating the pathways, proteins, and chemicals that facilitate
signaling in bacteria, to hundreds of groups that have integrated evolutionary biology,
computer science, mathematics, engineering, and metagenomics to create an ever-expanding
and dynamic field. In Quorum Sensing: Methods and Protocols, expert researchers provide
an in-depth set of diverse protocols that span this broad area of study.
Broken into three detailed sections, the volume covers the detection, isolation, and characterization of the QS signals made by both Gram- and Gram+ bacteria, determination of the function of QS signals in vivo, and the development of QS disruption strategies. Written in the highly successful Methods in Molecular Biology™ series format, chapters include brief introductions to their respective topics, lists of the necessary materials and reagents, step-by-step, readily reproducible laboratory protocols, and expert tips on troubleshooting and avoiding known experimental pitfalls. Comprehensive and cutting-edge, Quorum Sensing: Methods and Protocols serves as an invaluable collection of easily accessible techniques for scientists seeking to further our knowledge about bacterial communication and its relation to humanity.
Antibiofilm Agents: From Diagnosis to Treatment and Prevention
 This book provides a survey of recent advances in the development of antibiofilm agents
for clinical and environmental applications. The fact that microbes exist in structured
communities called biofilms has slowly become accepted within the medical community.
We now know that over 80% of all infectious diseases are biofilm-related; however,
significant challenges still lie in our ability to diagnose and treat these extremely
recalcitrant infections.
This book provides a survey of recent advances in the development of antibiofilm agents
for clinical and environmental applications. The fact that microbes exist in structured
communities called biofilms has slowly become accepted within the medical community.
We now know that over 80% of all infectious diseases are biofilm-related; however,
significant challenges still lie in our ability to diagnose and treat these extremely
recalcitrant infections.
Written by experts from around the globe, this book offers a valuable resource for medical professionals seeking to treat biofilm-related disease, academic and industry researchers interested in drug discovery, and instructors who teach courses on microbial pathogenesis and medical microbiology.
Hossain MA, Sattenapally N, Parikh HI, Li W, Rumbaugh KP, German NA. Design, synthesis, and evaluation of compounds capable of reducing Pseudomonas aeruginosa virulence. Eur J Med Chem. 2019 Oct 22:111800. PMID: 31706639
Tipton CD, Sanford NE, Everett JA, Gabrilska RA, Wolcott RD, Rumbaugh KP, Phillips CD. Chronic wound microbiome colonization on mouse model following cryogenic preservation. PLoS One. 2019 Aug 23;14(8):e0221565. PMID: 31442275
Zhao-Fleming HH, Wilkinson JE, Larumbe E, Dissanaike S, Rumbaugh K. Obligate anaerobes are abundant in human necrotizing soft tissue infection samples - a metagenomics analysis. APMIS. 2019 Aug;127(8):577-587. PMID: 31127652
Trivedi U, Madsen JS, Everett J, Fell C, Russel J, Haaber J, Crosby HA, Horswill AR, Burmølle M, Rumbaugh KP, Sørensen SJ. Staphylococcus aureus coagulases are exploitable yet stable public goods in clinically relevant conditions. Proc Natl Acad Sci U S A. 2018 Dec 11;115(50):E11771-E11779. doi: 10.1073/pnas.1804850115. Epub 2018 Nov 21.PMID:30463950
Zhao-Fleming H, Hand A, Zhang K, Polak R, Northcut A, Jacob D, Dissanaike S, and Rumbaugh KP. Effect of non-steroidal anti-inflammatory drugs on post-surgical complications against the backdrop of the opioid crisis. Burns & Trauma. 2018;6(1):25.
Winstanley C. and Rumbaugh K. Editorial: complexity and adaptability: an introduction to the special thematic issue on the genus Pseudomonas. FEMS Microbiol Lett. 2018 Sep 1:365(18). PMID: 30184124
Fleming D, Rumbaugh K. The Consequences of Biofilm Dispersal on the Host. Sci Rep. 2018 Jul 16;8(1):10738. PMID:30013112
Bjarnsholt T, Buhlin K, Dufrêne YF, Gomelsky M, Moroni A, Ramstedt M, Rumbaugh KP, Schulte T, Sun L, Åkerlund B, Römling U. Biofilm formation - What we can learn from recent developments. J Intern Med. 2018 Jun 1. doi: 10.1111/joim.12782. PMID: 29856510
Cornforth DM, Dees JL, Ibberson CB, Huse HK, Mathiesen IH, Kirketerp-Møller K, Wolcott RD, Rumbaugh KP, Bjarnsholt T, Whiteley M. Pseudomonas aeruginosa transcriptome during human infection. Proc Natl Acad Sci U S A. 2018 May 29;115(22):E5125-E5134. PMID: 29856510
Everett J, Gabrilska R, Rumbaugh KP, Vikström E. Assessing Pseudomonas aeruginosa Autoinducer Effects on Mammalian Epithelial Cells. Methods Mol Biol. 2018;1673:213-225. PMID: 29130176
Hamerly T, Everett JA, Paris N, Fisher ST, Karunamurthy A, James GA, Rumbaugh KP, Rhoads DD, Bothner B. Detection of Pseudomonas aeruginosa biomarkers from thermally injured mice in situ using imaging mass spectrometry. Anal Biochem. 2017 Oct 28;539:144-148. PMID:29107579
Smith A.C, Rice A., Sutton R.B., Gabrilska R., Wessel A.K., Whiteley M., and Rumbaugh K.P. Albumin inhibits Pseudomonas aeruginosa quorum sensing and alters polymicrobial interactions. Infect Immun. 2017 Jun 19. pii: IAI.00116-17. doi: 10.1128/IAI.00116-17. PMID: 28630071
C. B. Ibberson, A. Stacy, D. Fleming, J. L. Dees, M. S. Gilmore, K. Rumbaugh, and M. Whiteley. Co-infecting microorganisms dramatically alter pathogen gene essentiality
during polymicrobial infection. Nat Microbiol. 2017 May 30;2:17079. PMID:28555625
Zhao-Fleming H, Dissanaike S, Rumbaugh K. Are anaerobes a major, underappreciated cause of necrotizing infections? Anaerobe.
2017 Apr 24. PMID:28450145
Ahmad I., Khan M.S., Altaf, M.M., Qais, F.A., Ansari, F. A and Rumbaugh K.P. Biofilms: An overview of their significance in plant and soil health. Biofilms in Plant and Soil Health, First edition 2017. John Wiley &Sons, Ltd. Pp. 1-25.
Fleming D and Rumbaugh KP. Approaches to Dispersing Medical Biofilms. Microorganisms. 2017 Apr 1;5(2). PMID:28368320
Everett J, Turner K, Cai Q, Gordon V, Whiteley M, K. Rumbaugh. Arginine is a critical substrate for the pathogenesis of Pseudomonas aeruginosa in burn wound infections. MBio. 2017 Mar 14;8(2). PMID: 28292986
Fleming D, Chahin L, Rumbaugh K. Glycoside Hydrolases Degrade Polymicrobial Bacterial Biofilms in Wounds. Antimicrob Agents Chemother. 2017 Jan 24;61(2). PMID:27872074
Fleming D, Chahin L, Rumbaugh K. Glycoside Hydrolases Degrade Polymicrobial Bacterial Biofilms in Wounds. Antimicrob Agents Chemother. 2016 Nov 21. pii: AAC.01998-16. PMID:27872074
Trivedi U, Madsen JS, Rumbaugh KP, Wolcott RD, Burmølle M, Sørensen SJ. A post-planktonic era of in vitro infectious models: issues and changes addressed by a clinically relevant wound like media. Crit Rev Microbiol. 2016 Nov 21:1-13. PMID:27869519
Fleming D, Kesey J, Rumbaugh K, Dissanaike S. 2016. Comparing the Survivability of Lactobacillus Species in Various Probiotic Delivery Vehicles. JPEN J Parenter Enteral Nutr. 2016 Sep 29. PMID:27687914
Wolcott R, Sanford N, Gabrilska R, Oates JL, Wilkinson JE, Rumbaugh KP. 2016. Microbiota is a primary cause of pathogenesis of chronic wounds. J Wound Care. 2016 Oct;25(Sup10):S33-S43. PMID:27681809
M.M. Ramsey, M.O. Freire, R.A. Gabrilska, K.P. Rumbaugh, K.P. Lemon. Staphylococcus aureus shifts toward commensalism in response to Corynebacterium species. Front Microbiol. 2016 Aug 17;7:1230. PMID:27582729
C. Watters, D. Fleming, D.Bishop, K. P. Rumbaugh. Host Responses to Biofilm. Progress in Molecular Biology and Translational Science. 2016;142:193-239. PMID:27571696
A. Stacy, D. Fleming, R. J. Lamont, K. P. Rumbaugh, M. Whiteley. 2016. A commensal bacterium promotes virulence of an opportunistic pathogen via cross-respiration. MBio. June 28:7(3). PMID:27353758
Snow DE, Everett J, Mayer G, Cox SB, Miller B, Rumbaugh K, Wolcott RA, Wolcott RD. The presence of biofilm structures in atherosclerotic plaques of arteries from legs amputated as a complication of diabetic foot ulcers. J Wound Care. 2016 Feb;25(2):S16-22. PMID:26878370
Rai A, Pinto S, Velho TR, Ferreira AF, Moita C, Trivedi U, Evangelista M, Comune M, Rumbaugh KP, Simões PN, Moita L, Ferreira L. One-step synthesis of high-density peptide-conjugated gold nanoparticles with antimicrobial efficacy in a systemic infection model. Biomaterials. 2016 Apr;85:99-110. PMID:26866877
Gabrilska R and Rumbaugh KP. Biofilm Models of Polymicrobial Infection. Future Microbiol. 2015. Dec;10:1997-2015. PMID:26592098
Harrison F, Roberts AE, Gabrilska R, Rumbaugh KP, Lee C, Diggle SP. A 1,000-Year-Old Antimicrobial Remedy with Antistaphylococcal Activity. MBio. 2015 Aug 11;6(4). PMID: 26265721
Clinton A and Rumbaugh KP. Interspecies and Interkingdom Signaling via Quorum Signals. Israel Journal of Chemistry. 2015. 21 APR 2015.DOI: 10.1002/ijch.201400132
Watters C, Yuan TT, Rumbaugh KP. Beneficial and deleterious bacterial–host interactions in chronic wound pathophysiology. Chronic Wound Care Management and Research. 2015 April, Volume 2:53-62.
DeLeon S, Clinton A, Fowler H, Everett J, Horswill AR, Rumbaugh KP. Synergistic Interactions of Pseudomonas aeruginosa and Staphylococcus aureus in an In Vitro Wound Model. Infect Immun. 2014 Aug 25. pii: IAI.02198-14. PMID:25156721
Rumbaugh KP. Biofilms in the ICU. SWRCCC. 2014; 2(6):15-18. doi: 10.12746/swrccc2014.0206.067
Turner KH, Everett J, Trivedi U, Rumbaugh KP, Whiteley M. Requirements for Pseudomonas aeruginosa Acute Burn and Chronic Surgical Wound Infection. PLoS Genet. 2014 Jul 24;10(7):e1004518. PMID: 25057820
Trivedi U, Parameswaran S, Armstrong A, Burgueno-Vega D, Griswold J, Dissanaike S, Rumbaugh KP. Prevalence of Multiple Antibiotic Resistant Infections in Diabetic versus Nondiabetic Wounds. J Pathog. 2014;2014:173053. PMID: 25054067
Rumbaugh KP. Genomic complexity and plasticity ensure Pseudomonas success. FEMS Microbiol Lett. 2014 Jul;356(2):141-3. PMID: 25060810
Everett JA, Rumbaugh KP. “Biofilms, Quorum Sensing and Crosstalk in Medically-Important Microbes” (Chapter 14), in Tang Y, Liu D, Poxton IR, Schwartzman JD, and Sussman (eds.), “Molecular Medical Microbiology, 2nd Ed., 2014.
Rumbaugh KP and Armstrong A. “The Role of Quorum Sensing in Biofilm Development”, in Rumbaugh, K.P. and Ahmad, I. (eds). Antibiofilm Agents: From diagnosis to treatment and prevention (Springer Series on Biofilms) pgs.97-113. 2014. Springer, Humana Press, New York, NY. ISBN: 978-3-642-53832-2
Stacy A, Everett J, Jorth P, Trivedi U, Rumbaugh KP, Whiteley M. Bacterial fight-and-flight responses enhance virulence in a polymicrobial infection. Proc Natl Acad Sci U S A. 2014 May 13. pii: 201400586. [Epub ahead of print]. PMID: 24825893
Gawande PV, Clinton AP, LoVetri K, Yakandawala N, Rumbaugh KP, Madhyastha S. Antibiofilm Efficacy of DispersinB® Wound Spray Used in Combination with a Silver Wound Dressing. Microbiology Insights 2014:7, 9-13. PMID: 24826078
Watters C, Everett JA, Haley C, Clinton A, Rumbaugh KP. Insulin Treatment Modulates the Host Immune System To Enhance Pseudomonas aeruginosa Wound Biofilms. Infect Immun. 2014 Jan;82(1):92-100.
Jorth P, Trivedi U, Rumbaugh K, Whiteley M. Probing bacterial metabolism during infection using high-resolution transcriptomics. J Bacteriol. 2013 Aug 23. [Epub ahead of print]. PMID:23974023
Ilangovan A, Fletcher M, Rampioni G, Pustelny C, Rumbaugh K, Heeb S, Cámara M, Truman A, Chhabra SR, Emsley J, Williams P. Structural Basis for Native Agonist and Synthetic Inhibitor Recognition by the Pseudomonas aeruginosa Quorum Sensing Regulator PqsR (MvfR). PLoS Pathog. 2013 Jul;9(7):e1003508. PMID:23935486
Filiatrault MJ, Tombline G, Wagner VE, Van Alst N, Rumbaugh K, Sokol P, Schwingel J, Iglewski BH. 2013. Pseudomonas aeruginosa PA1006, Which Plays a Role in Molybdenum Homeostasis, Is Required for Nitrate Utilization, Biofilm Formation, and Virulence. PLoS One.8(2):e55594. doi: 10.1371/journal.pone.0055594. Epub 2013 Feb 8. PMID:23409004
A. Korgaonkar , U. Trivedi , K.P. Rumbaugh, M. Whiteley. 2013. Community surveillance enhances P. aeruginosa virulence during polymicrobial infection. Proc Natl Acad Sci U S A. 2013 Jan 15;110(3):1059-64. PMID:23277552
Watters C., Deleon K., Trivedi U., Griswold J.A., Lyte M., Hampel K.J., Wargo M.J., Rumbaugh K.P. 2012. Pseudomonas aeruginosa biofilms perturb wound resolution and antibiotic tolerance in diabetic mice. Med Microbiol Immunol. Sep 25. [Epub ahead of print]
Luckett, J.C.A, Darch, O., Watters, C., AbuOun, M., Ward, J., Goto, H., Heeb, S., Pommier, S., Rumbaugh, K., Camara, M., and Hardie, K.R. 2012. A novel virulence strategy for Pseudomonas aeruginosa mediated by an autotransporter with arginine-specific aminopeptidase activity. PLoS Pathogens. Aug;8(8):e1002854. PMID: 22927813
Padmanabhan V, Khan ZS, Solomon DE, Armstrong A, Rumbaugh KP, Vanapalli SA, Blawzdziewicz J. 2012. Locomotion of C. elegans: A Piecewise-Harmonic Curvature Representation of Nematode Behavior. PLoS One. 2012;7(7):e40121. PMID:22792224
Rumbaugh KP, Trivedi U, Watters C, Burton-Chellew MN, Diggle SP, West SA. 2012. Kin selection, quorum sensing and virulence in pathogenic bacteria. Proc Biol Sci. Sep 7;279(1742):3584-8. PMID:22648154
Stevens AM, Schuster M, Rumbaugh KP. 2012. Working together for the common good: cell-cell communication in bacteria. J Bacteriol. May;194(9):2131-41. PMID:22389476
Rumbaugh, K.P., Kaufmann, G.F. 2012. Exploitation of host signaling pathways by microbial quorum sensing signals. Curr Opin Microbiol. Apr;15(2):162-8. PMID: 22204809
Dalton, T., Dowd, S. E., Wolcott, R.D., Sun, Y., Watters, C., Griswold, J.A. and Rumbaugh, K.P. 2011. An in vivo polymicrobial biofilm wound infection model to study interspecies interactions. PLoS One. 6(11):e27317. PMID:22076151
Ramsey, M.C., Rumbaugh, K.P. and Whiteley, M. 2011. Metabolic cross-feeding enhances virulence in a model polymicrobial infection. PLoS Pathogens Mar;7(3):e1002012. PMID:21483753
Rumbaugh K.P. Fatal attraction: bacterial bait lures worms to their death. Proc Natl Acad Sci U S A. 2010 Sep 21;107(38):16411-2. Epub 2010 Sep 7.
Bryan, A., Koenig, L., Youn, E., Olmos, A., Li, G., Williams, S. C. and Rumbaugh, K.P. Human transcriptome analysis reveals a potential role for active transport in the metabolism of Pseudomonas aeruginosa autoinducers. Microbes Infect. 2010 Jul 24. [Epub ahead of print]
Teplitski, M., Mathesius, U. and Rumbaugh, K.P. Quorum sensing signal perception by mammalian and plant cells. Chem Rev. Chem Rev. 2010 Jun 10. [Epub ahead of print]
Rampioni, G., Pustelny, C., Fletcher, M.P., Wright, V. J., Bruce, M., Rumbaugh, K.P., Heeb, S., Camara, M., and Williams, P. 2010. Transcriptomic analysis reveals a global alkyl-quinolone-independent regulatory role for PqsE in facilitating the environmental adaptation of Pseudomonas aeruginosa to plant and animal hosts. Environ Microbiol. April 7. [Epub ahead of print]
Rumbaugh, K.P. and Carty, N. L. In vivo models of biofilm infection. In: Biofilm Infections. 2010. Springer, New York, NY. In press.
Jahoor, A., Williams, S.C., and Rumbaugh, K.P. Microbial signaling compounds as endocrine effectors. In: Microbial Endocrinology, Interkingdom Signaling in Infectious Disease and Health. 2010. Springer, New York, NY.
Wolcott, R.D.,Rumbaugh, K.P., James, G., Schultz, G., Phillips, P., Yang, Q., Watters, C., Stewart, P. S. and Dowd, S. E. 2010. Biofilm maturity studies indicate sharp debridement opens a time-dependent therapeutic window. J Wound Care 19(8) 320-328.
Rumbaugh, K.P., Diggle, S. P., Watters, C. W., Ross-Gillespie, A., Griffin, A. S. and West, S.A. 2009. Quorum sensing and the social evolution of bacterial virulence. Current Biology. Feb 19.
DeLeon, K., Watters, C., Baldin, F., Hamood, A., Griswold, J., Sreedharan, S., and Rumbaugh, K.P. 2009. Efficacy of gallium maltolate in treating Pseudomonas aeruginosa infection in a thermally-injured mouse model. Antimicrob Agents Chemother. Apr; 53(4):1331-1337.
Rumbaugh, K.P., 2009. Should we be afraid of the Green Monster? Crit Care Med. May;37(5):1826-7.
Jahoor A., Patel R., Bryan, A., Do C., Krier J., Watters C., Wahli W., Li, G., Williams S.C. ,Rumbaugh, K.P. 2008. Peroxisome Proliferator Activated Receptors Mediate Host Cell Pro-inflammatory Responses to P. aeruginosa Autoinducer. 2008. J Bacteriol. Jan 4.
Rumbaugh, K.P. 2007. Convergence of Hormones and Autoinducers at the Host/Pathogen Interface. Anal Bioanal Chem. 387(2): 425-435.
Schaber J.A., Triffo W.J., Suh S.J., Oliver J.W., Hastert M.C., Griswold J.A., Auer M., Hamood A.N., Rumbaugh, K.P. 2007. Pseudomonas aeruginosa forms Biofilms in Acute Infection Independently of Cell-to-Cell Signaling. Infect Immun. 75(8)p. 3715-21.
Shiner, E.K., Terentyev, D., Bryan, A., Sennoune, S., Martinez-Zaguilan, R., Li, G., Gyorke, S., Williams, S.C. and Rumbaugh, K.P. 2006. Pseudomonas aeruginosa Autoinducer Modulates Host Cell Responses through Calcium Signaling. Cellular Microbiol. 8(10):1601-10.
Beale, E., Li, G., Tan, M.W., Rumbaugh, K.P. 2006. Caenorhabditis elegans Senses Bacterial Autoinducers. Appl Environ Microbiol. 72(7):5135-7.
Shiner, E.K., Rumbaugh, K.P., and S. C. Williams. 2005. Interkingdom Signaling: deciphering the language of homoserine lactones. FEMS Microbiol Rev. 29(5):935-47.
Haynes, A., Ruda, F., Oliver, J., Hamood. A.N., Griswold, J.A., Park, P.W., Rumbaugh, K.P. 2005. Syndecan-1 Shedding Contributes to Pseudomonas aeruginosa Sepsis. Infect Immun. 73(12):7914-21.
Haynes, A., Rumbaugh, K.P., Park, W. P., Hamood, A. N., Griswold, J. A. 2005. Protamine sulfate reduces the susceptibility of thermally injured mice to Pseudomonas aeruginosa infection. J. Surg. Res. 123: 109-117.
Shiner, E.K., S. Reddy, C. Timmons, G. Li, S.C. Williams, and Rumbaugh, K.P. 2004. Construction of a bacterial autoinducer detection system in mammalian cells. Biol Proced Online. 6:268-276.
Williams, S.C. Patterson, E. K., Carty, N.L., Griswold, J. A., Hamood, A. N. and Rumbaugh, K.P. 2004. Pseudomonas aeruginosa autoinducer enters and functions in mammalian cells. J. Bacteriol. 186(8): 2281-7.
Rumbaugh, K.P. 2004. The Language of Bacteria…and Just About Everything Else. The Scientist. 18(17):26-27.
Rumbaugh, K.P., A. N. Hamood and J. A. Griswold. 2004. Cytokine Induction by the P. aeruginosa quorum sensing system during thermal injury. J Surg Res. 116:137-144.
Rumbaugh, K.P., J. A. Colmer, J. A. Griswold, and A. N. Hamood 2001. The effects of infection of thermal injury by Pseudomonas aeruginosa PAO1 on the murine cytokine response. Cytokine. 16:160-168.
Rumbaugh, K.P., J. A. Griswold, and A. N. Hamood 2000. The role of quorum sensing in the in vivo virulence of Pseudomonas aeruginosa. Microbes Infect. 2:1-11.
Rumbaugh, K.P., J. A. Griswold, B. H. Iglewski, and A. N. Hamood. 1999. Contribution of quorum sensing to the virulence of Pseudomonas aeruginosa in burn wound infection. Infect. Immun. 67:5854-5862.
Rumbaugh, K.P., J. Griswold, and A. Hamood. 1999. Contribution of the regulatory gene LasR to the pathogenesis of Pseudomonas aeruginosa infection of burned mice. J Burn Care Rehabil. 20:42-49.
Rumbaugh, K.P., J. A. Griswold, and A. N. Hamood. 1999. Pseudomonas aeruginosa strains obtained from patients with tracheal, urinary tract, and wound infection: variations in virulence factors and virulence gene. J. Hosp. Infect. 43:211-218.
Rumbaugh, K.P., A. N. Hamood, and J. A. Griswold. 1999. Analysis of Pseudomonas aeruginosa clinical isolates for possible variations within the virulence genes exotoxin A and exoenzyme S. J Surg Res. 82:95-105.
