Andrey L. Karamyshev, Ph.D.
Associate Professor

Ph.D. Biochemistry
Russian Academy of Sciences
Curriculum Vitae
Department of Cell Biology and Biochemistry
Texas Tech University Health Sciences Center
3601 4th Street, Lubbock, TX 79430-6540
Office Phone: (806) 743-4102
andrey.karamyshev@ttuhsc.edu
Research Interests
Molecular Mechanisms of Translational Regulation, RNA/Protein Quality Control and Protein Interactions in Health and Disease. Protein Synthesis, Targeting, Folding and Transport. Mechanisms of Protein Misfolding in Neurodegenerative Diseases.
Current Projects
Regulation of Aberrant Protein Production in Health and Disease.
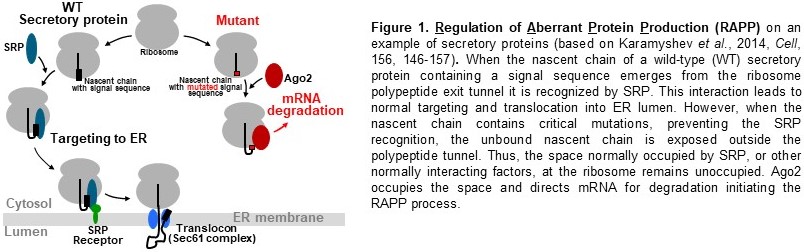
This project is based on our recent discovery of the novel pathway of translational
regulation and protein quality control known as RAPP (Regulation of Aberrant Protein
Production) (see Cell, 2014, 156, 146-157 and Frontiers in Genetics, 2018, 9, 431
for details) (Fig. 1). This pathway monitors the status of nascent chain interactions
during translation and transfers the signal to the mRNA degradation machinery to prevent
the synthesis of proteins that lost these natural interactions. During protein synthesis
the nascent chains that emerge from the ribosomal tunnel interact with multiple factors
for proper folding, targeting and modifications. These interactions are critical for
normal protein biogenesis. As we demonstrated, the synthesis of the aberrant secretory
proteins that are not able to interact with the targeting factor SRP (Signal Recognition
Particle) leads to the activation of the RAPP pathway and the destruction of the defective
protein’s mRNA. While the existence of the RAPP pathway is well documented, its mechanism
is practically unknown. Currently, we identified only one component of the RAPP pathway,
AGO2. AGO2 is a key element of the RAPP process and it triggers mRNA degradation.
There are many questions that remain to be answered. What are the other components
of the RAPP machinery? What are signals for mRNA degradation? Has mRNA cleavage happened
at the ribosome? Which enzyme cleaves mRNA? How is the signal about aberrant nascent
chain transmitted to mRNA degradation machinery? To answer these questions we use
complex approaches as in vivo (cultured human cells, immunofluorescence, microscopy,
RNAi, overexpression, etc.), as well as in vitro (protein translation systems, incorporation
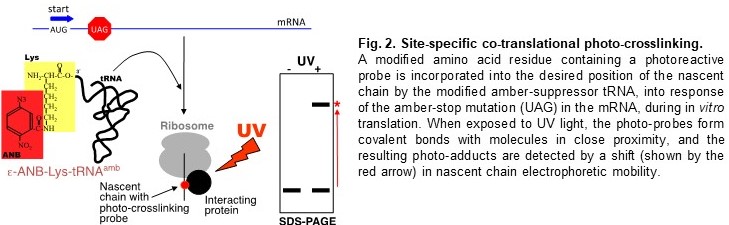
of unnatural amino acids by suppressor tRNAs, site-specific photo-crosslinking (Fig.
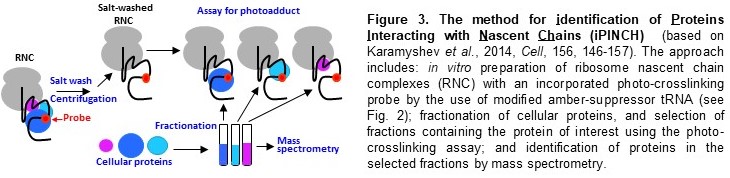
2), method for identification of Proteins Interacting with Nascent Chains (iPINCH)
(Fig. 3), and many others). The future directions of our research are focused on the
elucidation of the RAPP mechanism, its involvement in mRNA/protein quality control
of different types of aberrant proteins, and the role of RAPP in human diseases. Our
data demonstrate that pathological RAPP activation is a molecular basis for various
human diseases. It opens opportunity for identification of new pharmacological targets
and for the disease treatments.
Mechanisms of Protein Misfolding in Neurodegenerative Diseases.
Numerous neurodegenerative diseases are associated with the accumulation of protein
aggregates and the death of neurons. Alzheimer’s, Parkinson’s and Huntington’s diseases,
amyotrophic lateral sclerosis and others are caused by the aggregation of aberrant
proteins. This project takes our view at these proteins during the first steps in
their synthesis. Normally, during translation, the proteins interact with their partners
that assist in correct folding. We propose that the loss of these early interactions
leads to protein aggregation and disease. We use site-specific photo-crosslinking
(Fig. 2) and recently developed iPINCH (Fig. 3) to identify nascent chain interactions
during translation in health and in neurodegenerative diseases. This project is directed
at establishing a new and currently under-examined area of neurodegenerative research
— co-translational interactions of proteins and their roles in the disease state.
The interacting partners may serve as modulators of protein folding and aggregation.
Potentially this project will lead to the discoveries of novel therapeutics for the
treatment of neurodegenerative disease.
 |
 |
 |
Graduate students are welcome to the lab. Please contact Dr. Andrey Karamyshev if
you have an interest to join the lab.
Selected Publications
(From 54 publications)
- Gutierrez Guarnizo, S. A., Kellogg, M. K., Miller, S. C., Tikhonova, E. B., Karamysheva, Z. N. Karamyshev, A. L. (2023) Pathogenic Signal Peptide Variants in the Human Genome. NAR Genomics and Bioinformatics, 5(4). https://doi.org/10.1093/nargab/lqad093
- Orobets, K. S., and Karamyshev, A. L. (2023) Amyloid Precursor Protein and Alzheimer’s Disease. International Journal of Molecular Sciences, 24(19), 14794; PMID: 37834241. PMCID: PMC10573485. https://doi.org/10.3390/ijms241914794
- Karamysheva, Z. N., Karamyshev, A. L. (2023) Aberrant Protein Targeting Activates Quality Control on the Ribosome. Frontiers in Cell and Developmental Biology, 11: 1198184. PMID: 37346176. PMCID: PMC10279951. https://doi.org/10.3389/fcell.2023.1198184 (* = corresponding authors)
-
Gutierrez Guarnizo, S. A., Tikhonova, E. B., Karamyshev*, A. L., Muckus*, C. E., Karamysheva*, Z. N. (2023) Translational reprogramming as a driver of antimony-drug resistance in Leishmania. Nature Communications, 14, 2605. PMID: 37147291. PMCID: PMC10163012. https://doi.org/10.1038/s41467-023-38221-1. (* = corresponding authors).
The article was featured in:
Texas Tech Today. Finding Possible Solutions Through Working With Parasites by Doug Hensley. https://today.ttu.edu/posts/2023/06/Stories/Finding-Possible-Solutions-Through-Working-With-Parasites
EurekAlert! AAAS TTUHSC-TTU research collaboration leads to possible drug targets for Leishmaniasis: https://www.eurekalert.org/news-releases/993169
TTUHSC Daily Dose by Mark Hendricks. TTUHSC-TTU Research Collaboration Leads to Possible Drug Targets for Leishmaniasis (by Mark Hendricks). https://dailydose.ttuhsc.edu/2023/june/research-collaboration-Leishmaniasis.aspx
Editors’ Highlights by Madlen Luckner in Microbiology and Infectious Diseases, Nature Communications - Miller, S. C.; MacDonald, C. C., Kellogg, M. K., Karamysheva, Z. N., Karamyshev, A. L. (2023) Specialized Ribosomes in Health and Disease. International Journal of Molecular Sciences. 24(7): 6334. PMID: 37047306. PMCID: PMC10093926. https://doi.org/10.3390/ijms24076334.
- Rodríguez-Almonacid, C. C., Kellogg, M. K., Karamyshev, A. L., Karamysheva, Z. N. (2023) Ribosome Specialization in Protozoa Parasites. International Journal of Molecular Sciences, 24(8):7484. PMID: 37108644. PMCID: PMC10138883. https://doi.org/10.3390/ijms24087484.
- Tikhonova, E. B., Gutierrez Guarnizo, S. A., Kellogg, M. K., Karamyshev, A., Dozmorov, I. M., Karamysheva, Z. N., Karamyshev, A. L. (2022) Defective Human SRP Induces Protein Quality Control and Triggers Stress Response. Journal of Molecular Biology. vol. 434, Issue 22. PMID: 36210597. https://doi.org/10.1016/j.jmb.2022.167832
- Kellogg, M. K.; Tikhonova, E. B.; Karamyshev, A. L. (2022) Signal Recognition Particle in Human Diseases. Frontiers in Genetics, 13:898083, p. 1-7. https://doi.org/10.3389/fgene.2022.898083
- Hernandez, S. M.; Tikhonova, E. B.; Baca, K. R.; Zhao, F.; Zhu, X.; Karamyshev, A. L. (2021) Unexpected Implication of SRP and AGO2 in Parkinson’s Disease: Involvement in Alpha-Synuclein Biogenesis. Cells, 10, 2792. https://doi.org/10.3390/cells10102792
- Kellogg, M. K.; Miller, S. C.; Tikhonova, E. B.; Karamyshev, A. L. (2021) SRPassing Co-translational Targeting: The Role of the Signal Recognition Particle in Protein Targeting and mRNA Protection. International Journal of Molecular Sciences, 22, 6284. https://doi.org/10.3390/ijms22126284 PMID: 34208095 PMCID: PMC8230904
- Karamysheva, Z. N.; Moitra, S.; Perez, A.; Mukherjee, S.; Tikhonova, E. B.; Karamyshev, A. L.; Zhang, K. (2021) Unexpected Role of Sterol Synthesis in RNA Stability and Translation in Leishmania. Biomedicines, 9, 696. https://doi.org/10.3390/biomedicines9060696 PMID: 34205466 PMCID: PMC8235615
- Gutierrez Guarnizo, S. A., Tikhonova, E. B., Masoud Zabet-Moghaddam, Zhang, K., Muskus, C., Karamyshev A. L., Karamysheva, Z. N. (2021) Drug-Induced Lipid Remodeling in Leishmania Parasites. Microorganisms. 9, 790. https:// doi.org/10.3390/microorganisms 9040790.
- Lee, A. K., Klein, J., Fon Tacer, K., Lord, T., Oatley, M. J., Oatley, J. M., Porter, S. N., Pruett-Miller, S. M., Tikhonova, E. B., Karamyshev, A. L., Wang, Y. D., Yang, P., Korff, A., Kim, H. J., Taylor, J. P., Potts, P. R. (2020). Translational Repression of G3BP in Cancer and Germ Cells Suppresses Stress Granules and Enhances Stress Tolerance. Molecular Cell. 79: 645-659, doi: 10.1016/j.molcel.2020.06.037. PMID: 32692974.
- Karamyshev, A. L., Tikhonova, E. B., Karamysheva, Z. N. (2020) Translational Control of Secretory Proteins in Health and Disease. International Journal of Molecular Sciences. 21, 2538; doi:10.3390/ijms21072538. PubMed PMID: 32268488; PubMed Central PMCID: PMC7177344.
- Karamysheva, Z. N., Gutierrez Guarnizo, S. A., Karamyshev, A. L. (2020) Regulation of Translation in the Protozoan Parasite Leishmania. International Journal of Molecular Sciences. 2020, 21, 2981; doi:10.3390/ijms21082981. PubMed PMID: 32340274.
- Hernandez, S. M., Tikhonova, E. B., Karamyshev, A. L. (2020) Protein-Protein Interactions in Alpha-Synuclein Biogenesis: New Potential Targets in Parkinson’s Disease. Frontiers in Aging Neuroscience. 12:72; doi: 10.3389/fnagi.2020.00072. PubMed PMID: 32256340; PubMed Central PMCID: PMC7092629.
- Wang, D., Wang, T., Gill, A., Hilliard, T., Chen, F., Karamyshev, A. L., Zhang, F. (2020) Uncovering the Cellular Capacity for Intensive and Specific Feedback Self-control of the Argonautes and MicroRNA Targeting Activity. Nucleic Acids Research, 48, 9, 4681-4697. doi: 10.1093/nar/gkaa209. PubMed PMID: 32297952.
- Wang, D., Karamyshev, A. L. (2020) Next Generation Sequencing (NGS) Application in Multiparameter Gene Expression Analysis. Molecular Toxicology Protocols. Methods in Molecular Biology, vol. 2102, Springer. 17-34. PMID: 31989548, DOI: 10.1007/978-1-0716-0223-2_2
- Tikhonova, E. B., Karamysheva, Z. N., von Heijne, G., Karamyshev, A. L. (2019) Silencing of aberrant secretory protein expression by disease-associated mutations. Journal of Molecular Biology, vol. 431, p. 2567–2580. PMID: 31100385, doi: 10.1016/j.jmb.2019.05.011.
- Karamysheva, Z. N., Tikhonova, E. B., Karamyshev, A. L. (2019) Granulin in Frontotemporal Lobar Degeneration: Molecular Mechanisms of the Disease. Frontiers in Neuroscience, 13:395. doi: 10.3389/fnins.2019.00395. PMID: 31105517 PMCID: PMC6494926.
- Karamyshev, A. L., and Karamysheva, Z. N. (2018) Lost in Translation: Ribosome-Associated mRNA and Protein Quality Controls. Frontiers in Genetics, 9:431. doi: 10.3389/fgene.2018.00431. The article is in the top 5.0 % most viewed and downloaded articles in the 4 quarter of 2018 (Frontiers)
- Pinarbasi, E. S., Karamyshev, A. L., Tikhonova, E. B., Wu, I-H., Hudson, H., Thomas, P. J. (2018) Pathogenic signal sequence mutations in progranulin disrupt SRP interactions required for mRNA stability. Cell Reports, 23, 2844-2851, https://doi.org/10.1016/j.celrep.2018.05.003
- Karamysheva, Z. N., Tikhonova, E. B., Grozdanov, P. N., Huffman J. C., Baca, K. R., Karamyshev, A, Denison R. B., MacDonald, C. C., Zhang, K., Karamyshev, A. L. (2018) Polysome Profiling in Leishmania, Human Cells and Mouse Testis. Journal of Visualized Experiments (JoVE), 134, e57600, doi:10.3791/57600.
- Vetter,* A. J., Karamyshev,* A. L., Patrick, A. E., Hudson, H., Thomas, P. J. (2016) N-alpha-acetyltransferases and regulation of CFTR expression. PLoS ONE 11(5): e0155430. *These authors contributed equally to this work.
- Nilsson, I., Lara, P., Hessa T., Johnson, A. E., von Heijne, G., Karamyshev, A. L. (2015) The code for directing proteins for translocation across ER membrane: SRP cotranslationally recognizes specific features of a signal sequence. Journal of Molecular Biology, 427, 1191-1201.
-
Karamyshev,* A. L., Patrick, A. E., Karamysheva, Z. N., Griesemer, D. S., Hudson, H., Tjon-Kon-Sang, S, Nilsson, I., Otto, H., Liu, Q., Rospert, S., von Heijne, G., Johnson, A. E., Thomas,* P. J. (2014) Inefficient SRP Interaction with a Nascent Chain Triggers a mRNA Quality Control Pathway. Cell, 156, 146-157. (* = co-corresponding authors).
This paper was featured in:
Trends in Biochemical Sciences (2014), 39(4), 154-156. Defective secretory-protein mRNAs take the RAPP by Popp, M. W.-L. and Maquat L. E.
Nature Reviews Genetics (2014), 15(3), 144. Research highlights. Novel mRNA quality control mechanism.
Recommended by Faculty of 1000 - selected for F1000Prime. - Patrick A. E., Karamyshev A. L., Millen L., Thomas P. J. (2011) Alteration of CFTR transmembrane span integration by disease-causing mutations. Mol. Biol. Cell, 22(23), 4461-4471.
- Karamyshev, A. L., Kelleher, D. J., Gilmore, R., Johnson, A. E., von Heijne, G., Nilsson, I. (2005) Mapping the interaction of the STT3 subunit of the oligosaccharyl transferase complex with nascent polypeptide chains. J. Biol. Chem., 280, 49, 40489-40493.
- Karamyshev, A. L., Johnson, A. E. (2005) Selective SecA association with signal sequences in ribosome-bound nascent chains: A potential role for SecA in ribosome targeting to the bacterial membrane. J. Biol. Chem., 280, 45, 37930-37940. The article has been recommended by Faculty of 1000 - selected for F1000Prime.
- Karamyshev, A. L., Karamysheva, Z. N., Yamami, T., Ito, K., Nakamura, Y. (2004) Transient idling of posttermination ribosomes ready to reinitiate protein synthesis. Biochimie, 86(12), 933-938.
- Karamysheva, Z. N., Karamyshev, A. L., Ito, K., Yokogawa, T., Nishikawa, K., Nakamura, Y., Matsufuji, S. (2003) Antizyme frameshifting as a functional probe of eukaryotic translational termination. Nucleic Acids Res., 31(20), 5949-5956. The article has been recommended by Faculty of 1000 - selected for F1000Prime.
- Ito, K., Frolova, L., Seit-Nebi, A., Karamyshev, A. L., Kisselev, L., Nakamura, Y. (2002) Omnipotent decoding potential resides in eukaryotic translation termination factor eRF1 of variant-code organisms and is modulated by the interactions of amino acid sequences within domain 1. Proc. Natl. Acad. Sci. USA, 99(13), 8494-8499.
- Kervestin, S., Garnier, O. A., Karamyshev, A. L., Ito, K., Nakamura, Y., Meyer, E., Jean-Jean, O. (2002) Isolation and expression of two genes encoding eukaryotic release factor 1 from Paramecium tetraurelia. J. Eukaryot. Microbiol., 49(5), 374-382.
- Karamyshev, A. L., Ito, K., Nakamura, Y. (1999) Polypeptide release factor eRF1 from Tetrahymena thermophila: cDNA cloning, purification and complex formation with yeast eRF3. FEBS Lett., 457(3), 483-488.
- Karamyshev, A. L., Karamysheva, Z. N., Kajava, A. V., Ksenzenko, V. N., Nesmeyanova, M. A. (1998) Processing of Escherichia coli alkaline phosphatase: role of the primary structure of the signal peptide cleavage region. Journal of Molecular Biology, 277(4), 859-870.
